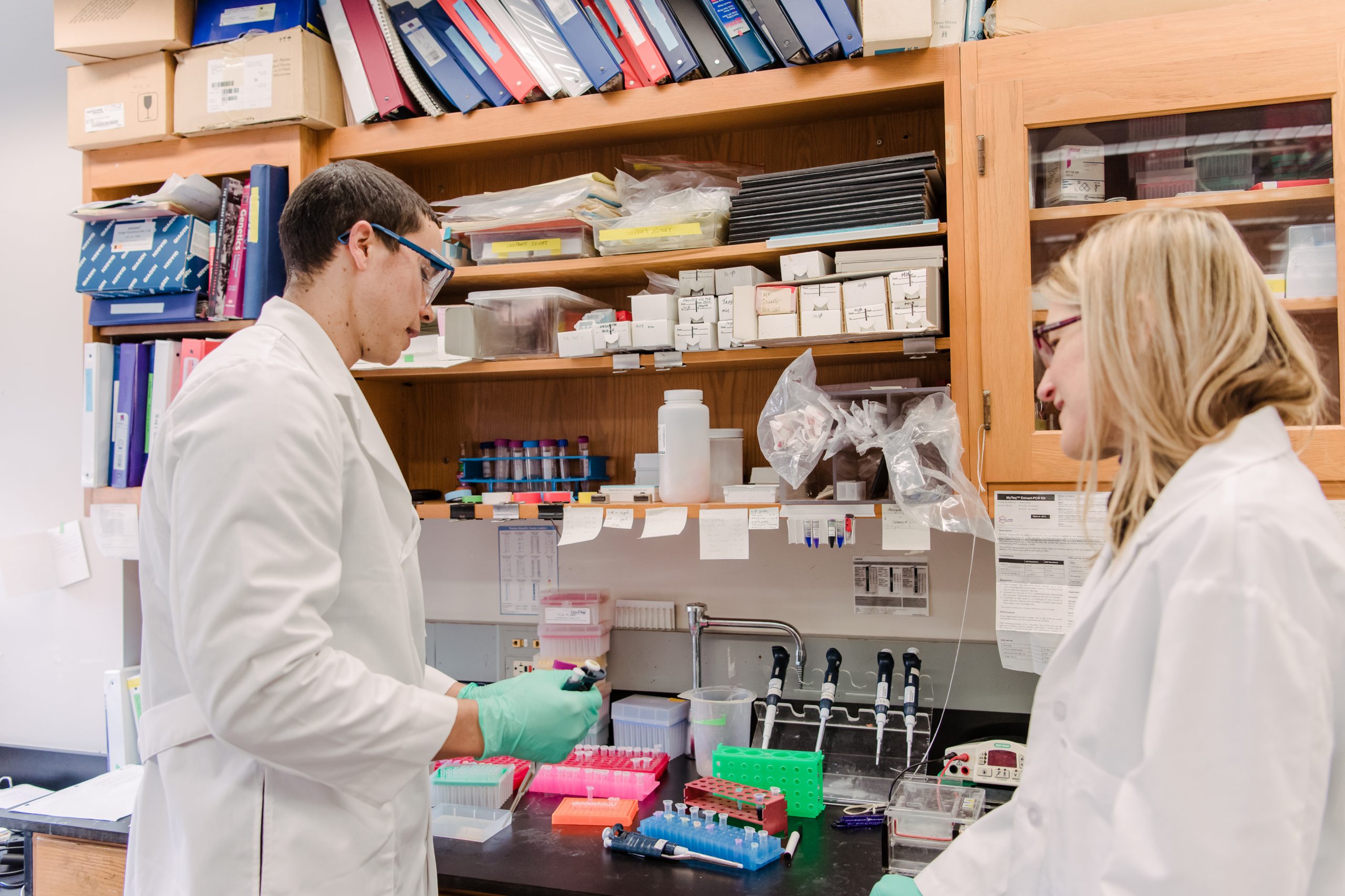
Cell migration—how, when, and why cells move—has important implications for understanding development and diseases such as blood cell disorders, rheumatoid arthritis, and metastatic cancer. Michelle Starz-Gaiano, associate professor of biological sciences, has learned a great deal about cell migration from observing Drosophila melanogaster, the humble fruit fly, in her lab. She’s also learned that experimental tools have their limitations.
“We have fantastic genetic tools in Drosophila, and we have great live imaging, so we can get pretty far doing that,” Starz-Gaiano says. “But then we encounter things that we can’t explain.”
That’s where her decade-plus collaboration with Brad Peercy, associate professor of mathematics, comes in. In their partnership, Peercy develops mathematical models to represent the movement of cells across developing eggs in the fly ovary. Now, a three-year, $370,000 NSF grant will support the duo as they combine their expertise to further explore the regulation of cell movement.
“I think this project has really big implications for how we think about development and how organs and tissues function,” Starz-Gaiano says, “because it’s looking at aspects that people haven’t yet paid attention to.”
A fresh perspective
Scientists can learn a lot about cell migration through experimental approaches like those Starz-Gaiano employs. However, some experiments are too complex or too expensive to attempt without a powerful reason to believe they will reveal new and useful information.
Turning to mathematical modeling has been particularly helpful for pointing wet lab scientists in the right direction before they invest in complicated experiments. “The collaboration has been incredible in making the best predictions for how to explain confusing results. It’s enabled us to narrow down the set of things to test,” Starz-Gaiano says.

For example, cell migration in the fruit fly ovary happens in several complex stages. Chemical signaling and the geometry of the ovary both play a role in regulating the process. “There are lots of different interesting features that are ripe for having a mathematical framework put around them,” Peercy says, “and the math can sometimes point to looking at something a little bit differently than biologists might otherwise.”
Now the challenge is that the math has pointed to certain attributes of the migration process that may be impossible to test in a live organism with today’s technology. “The models that Brad’s group has made are so compelling that now we have to figure out if we can show that biologically,” Starz-Gaiano says. “That’s a big focus of some of the work in the lab now.”
Uncharted territory
This new grant focuses on the math side of the work. Peercy and Starz-Gaiano have been collaborating since 2008, when they partnered to work with undergraduates as part of the former, NSF-funded Interdisciplinary Training for Undergraduates in Biological and Mathematical Sciences program.
Since then, Peercy and his team have developed several models for different aspects of the cell migration process. The next step is “integrating some of the models that we’ve already developed into a more comprehensive model of the system,” he explains. For that, Peercy and Starz-Gaiano will benefit from the computational expertise of Matthias Gobbert, professor of mathematics, who is another co-investigator on the grant. He will help make the computational processing more efficient, and therefore feasible in a reasonable amount of time.
Another aim of the project is to further investigate a set of chemical reactions, known as a signaling cascade, involved in triggering cell migration. “The signaling cascade that the model is for is very well conserved across species, and it’s implicated in a lot of human diseases,” Starz-Gaiano says. “The model pointed to a certain kind of regulation that hasn’t been very well explored, so we want to follow up on that.”

The team will also work to better understand how the restricted space of the egg chamber affects how cells and chemical signals move, and how that affects cell migration. “Being able to map that complicated geometry into a computational framework is something we’re interested in,” Peercy says. For this part of the project, they’re collaborating with Tagide deCarvalho, director of UMBC’s Keith R. Porter Imaging Facility.
Looking at the space between cells is a new area of research. “People haven’t done that,” Starz-Gaiano says. “They just think this one sends a signal and that one gets it and it’s over, and we’re saying, well, what happens in between?” Failure of a signal to reach its final destination can contribute to some birth defects, Starz-Gaiano explains. She also wonders if the same could be true for some cancers.
A new approach
As scientific discovery marches onward, researchers in fields that were once seemingly disparate find themselves relying on each other more and more. “The biology is insufficient to capture what’s going on, so we need different approaches,” Starz-Gaiano says.
Collaborations between researchers in different fields can be tricky to navigate, as each side learns to speak a new language and engage in different ways of thinking. But when everyone involved is on board, the results can be groundbreaking.
“You see strong collaborations between experimentalists and theoreticians when they are willing to sit in uncomfortable situations,” Peercy says. That might mean a biologist thinking about differential equations or a mathematician trying to understand the reproductive process of a fruit fly.
Peercy and Starz-Gaiano’s easy rapport and long history of working together makes it clear that they’ve crossed that bridge and are comfortable taking on the unknown together. They recognize the benefits that combining their unique research skill sets can offer.
“The only way that you can solve complex problems now is to use multiple strategies at the same time,” Starz-Gaiano says. “And increasingly, that’s what we’re faced with in biology—these problems are too complicated for one method to tackle them.”
Banner image: Michelle Starz-Gaiano (right) works with Jeffrey Inen ’18 in her lab. Photo by Marlayna Demond ’11 for UMBC.