Like bacteria, fungi can cause disease inside your body and on your skin, or even grow on medical equipment like catheter tubing and wound dressings. Many fungal diseases are treatable with antifungal medications, but drug resistance is a growing problem. With a new four-year, $1.3 million grant from NIH, Jeffrey Gardner and his students will be looking for new ways to target disease-causing fungi.
Typically, drugs that treat fungal disease prevent the fungus from making its cell wall, which either kills the fungus outright or weakens it enough that the immune system can finish it off. But that method is highly susceptible to developing resistance.
“But what if you had an external attack on the fungus? What if there were enzymes that actively degraded the fungal cell wall, which is not going to generate resistance easily?” asks Gardner, associate professor of biological sciences. “Our goal is to find enzymes that effectively break down the fungal cell wall, that can be used as a treatment in parallel with cell wall synthesis blockers, or on their own.”
Homing in on the right enzyme
With the new support from NIH, Gardner’s lab will investigate hundreds of bacterial enzymes to figure out which best target Aspergillus fungus, a group of common disease-causing mold species. Previous work from other groups has found particular bacterial species that slow down fungal growth. Gardner’s team will start by looking at every gene and protein in these bacteria to figure out “which genes are turned on, and which proteins are made, while the bacteria are degrading and eating the fungus,” he explains.
Gardner expects that should whittle down the candidate list from about 4,000 genes to a few hundred. “How do we figure out which ones of those couple hundred actually matter? That’s where our genetic system comes in,” he says. The team will generate bacterial strains that lack a functional version of each of the candidate genes and see which strains are less efficient at eating the fungus.
That process should further narrow the list of genes to a few dozen, Gardner says. Then, they’ll produce strains that are missing different combinations of the candidate genes. When they find a combination that can’t survive at all on fungus alone, they’ll be confident the genes knocked out in that strain are the ones necessary to eat the fungus. Finally, they’ll run biochemistry experiments to determine the function and mechanism of each enzyme.

From one to a billion
This kind of brute-force methodology, involving hundreds of unique strains of bacteria, highlights the advantages of working with species that multiply incredibly fast. “That’s the power of microbiology—you can go from a single cell to an overnight culture with a billion cells,” Gardner says. “You can do many, many experiments at a scale and at a speed that really isn’t possible with, say, a fly or a mouse system.”
The culmination of the project will be “identifying the enzymes, knowing what they’re doing, how they’re doing it, and how well they’re doing it,” Gardner says.
The power of microbes
As a microbial researcher who cares about “interesting bits of biology that have no medical relevance at all,” Gardner says he never imagined he would receive an NIH grant to pursue work with clear biomedical implications. But as his research at UMBC over the last decade has progressed, he’s realized his work can transcend typical disciplinary boundaries. “I think that’s the exciting part, and the power of microbial systems, that you have that latitude,” Gardner says.
His first major grant from the Department of Energy focused on bacteria’s role in biofuel production. A second project funded by the National Science Foundation looks at how bacteria contribute to the carbon cycle by breaking down dead matter on the forest floor. And now, with funding from NIH, his group will look at how bacteria can help fight fungal disease.
“If you can find an interesting bug, with some interesting physiology, the types of questions can really span major different areas,” Gardner says.
The techniques often remain the same, though. “Our wheelhouse is genetics, systems biology, and physiology,” Gardner says. In the new project, “We’re leveraging an established pipeline to work on a problem that is a step away from what we have traditionally done. We’ve got lots of tools and tricks for working with the bacteria.”
The wide range of methods used in Gardner’s lab means that his group offers great training opportunities for students. With the new grant, Gardner is looking forward to adding two new Ph.D. students and several undergraduate researchers to his team.
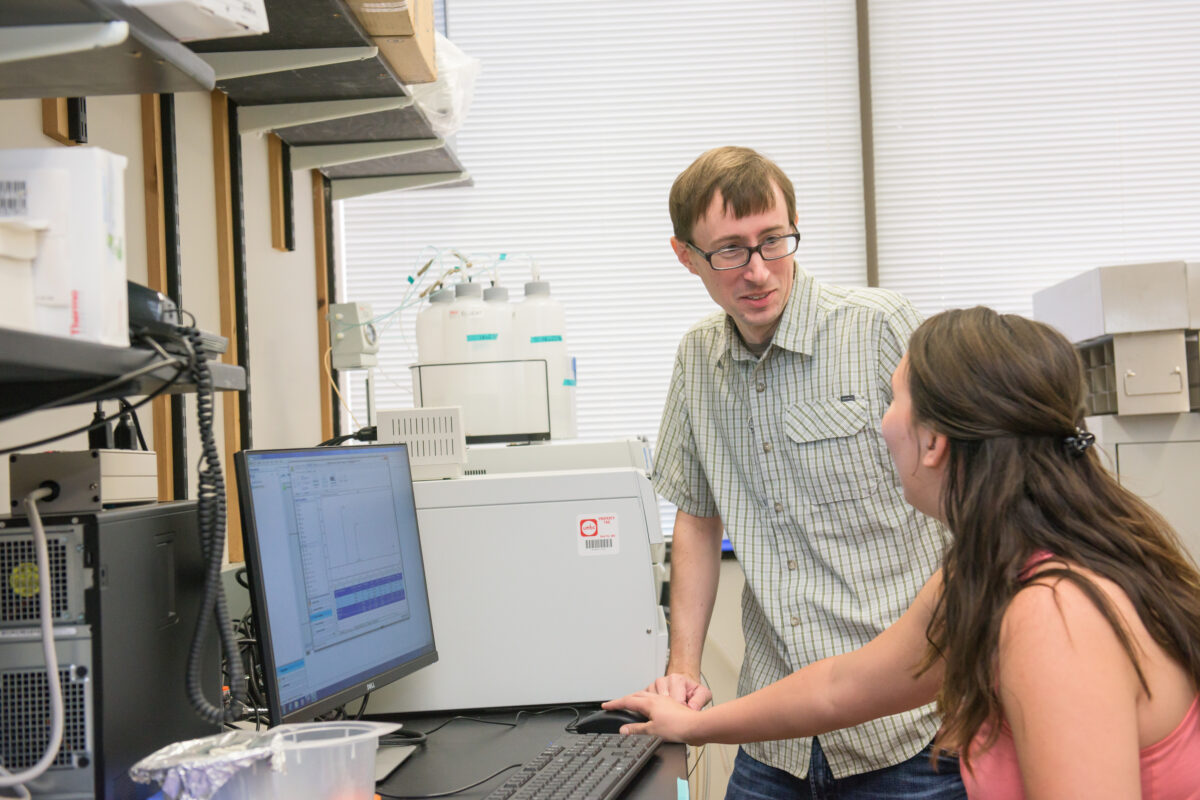

Essential support
Gardner knows, from personal experience, the importance of research training and mentorship along a scientist’s full career path, including in his time at UMBC. Before applying for this current grant, he took advantage of a program offered by UMBC’s College of Natural and Mathematical Sciences specifically for people seeking major NIH grants. The program, orchestrated by Phyllis Robinson, professor of biological sciences, included workshops, critique partners, and mentoring. “That was absolutely essential for my success,” Gardner says.
Moving forward, if the project goes well and Gardner is able to secure a second round of funding in a few years, he says the next questions will be, “How do we engineer the enzymes to be more potent? How do we make them do their job of killing fungi better?” The lab could even ask which combination of enzymes would target a specific fungus most effectively, or provide the broadest protection against many fungal species.
But for now, they’ll focus on nailing down the basics, he says. “We need to find out who’s there and what they’re doing first.”